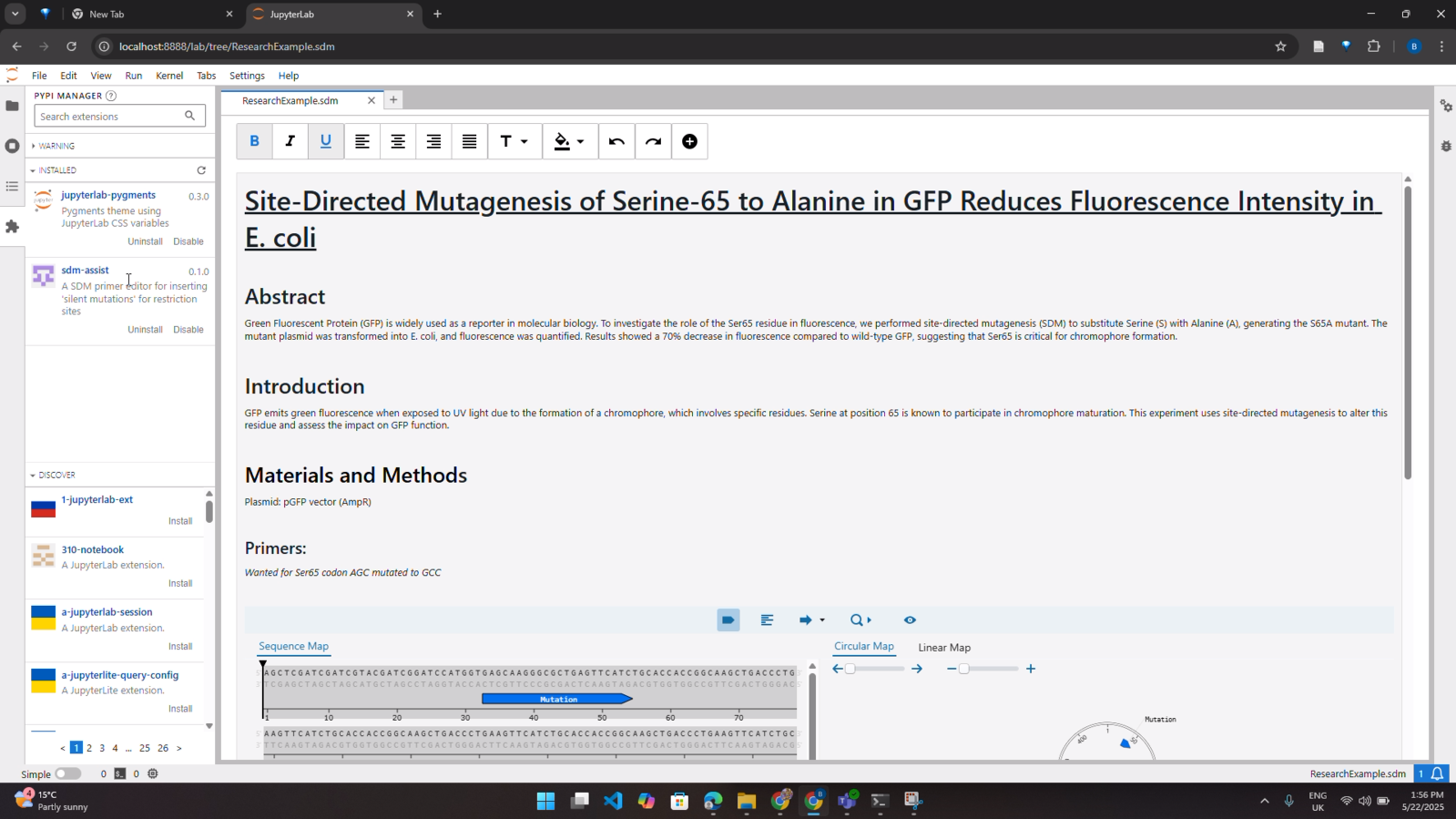
Overview
Redesigned and rebuilt a legacy molecular biology tool used to support site-directed mutagenesis workflows (SDM Assist) from an ageing ActionScript 3 and Adobe AIR stack into a modern React and TypeScript application running as a custom JupyterLab extension.
What it does
SDM-Assist helps researchers design mutagenic primers and identify silent restriction-site changes that can be used to screen for successful mutations.
Users can:
- Import DNA sequences from a variety of formats (including FASTA, Genbank & raw ATCG)
- View in visualiser and switch views between nucleotide and amino-acid representations
- Select codons and visually insert mutations
- Select type of mutation:
- Change to specified codon of a specified amino acid
- Let algorithm choose best codon for specified amino acid
- Generate and export scored primer candidates with supporting statistics
- Compare mutation and primer candidates
- Write structured, stylable experiment notes around embedded primers and sequence visualisations
Why it’s interesting
It was a multi-layer software engineering project spanning interface design, document architecture, state handling and domain-specific algorithm porting. I had to understand a specialist legacy codebase, preserve a familiar workflow for existing users and translate core biological logic into a maintainable modern platform. I also added some new quality of life and workflow improvement features such as undo/redo, better visualisations and storage for experiment notes.
Architecture
The project was designed as a multi-layer application rather than just a frontend because SDM-Assist needed to combine an interactive interface, custom document handling and domain-specific biological logic into one coherent workflow.
- Presentation Layer: Built with Material UI and custom React components including: dialogs, result tables and editor controls. A customised Slate.js served as the rich text editor for experiment notes and a modified TeselaGen Open Vector Editor (OVE) acted as the sequence visualiser.
- Application: Local React state and Redux orchestrated the editing workflow, undo/redo and composition of UI pieces. The modified TeselaGen OVE component interfaced directly with business logic to insert mutations, generate, score and display candidate primers.
- Business-Logic/Domain Layer: TypeScript implementations of mutagenesis, primer-generation and scoring algorithms that embody the core biological rules.
- Platform & Persistence Layers: JupyterLab served as the project’s platform. Project was integrated with a custom document type, document model and widget as well as all of their respective factories. Output files were serialised to a bespoke .sdm file format with state, experiment notes, primer/sequence and metadata storage allowing my project to act like a first-class document within JupyterLab rather than being a loose embedded panel.
Key Technical Points
- Custom JupyterLab Architecture: Built as a custom JupyterLab extension with its own file type that heavily extended the JupyterLab Document Model and Registry. Enabling users to maintain saved and partial states during the primer design process without ever losing their progress.
- Local Processing: All sequence and domain/business logic is run locally because researchers can’t always upload sequence data to third-party tools as they may be proprietary or private.
- Workflow Improvements: Added many workflow improvements such as: global undo/redo, document save-and-resume and the ability to attach experiment notes to designed primers.
- Advanced Visualisations: Adapted the TeselaGen OVE component for DNA and protein visualisation, mutation selection and insertion + interactive sequence editing within the document.
- Ported from existing codebase: Translated the core mutagenesis, primer-generation and scoring logic from the original AS3 codebase into modular, maintainable TypeScript while refactoring the OOP design to improve maintainability and readability.
Tech Stack
Frontend / Platform: React, TypeScript, JupyterLab
Bioinformatics Tooling: TeselaGen OVE, bio_parsers & sequence-utils libraries
UI / Other libs: Material UI, Slate.js, Redux
Limitations
The project successfully reproduced the complex mutagenesis workflow and much of the original tool’s behaviour, though a minor bug in the Tm calculation logic remains as an area for future refinement.